Diagnostic therapy efficacy assessment as a novel procedure in genomic medicine and rare cancer care.
Precision medicine in cancer care aims to target the treatment of an individual patient on basis of a tailored diagnosis. The advent of genomic big data describing the genetic landscape of human cancers has revived the concept of oncogene defined molecular cancer grouping and resulted in genomic medicine. International efforts providing a pan-cancer comparison of the genomic similarities of tens of thousands of patients now suggest shifting the grouping of cancers towards tissue-agnostic subtypes defined by the genomic aberrations rather than the tissue or cell of origin. This has a profound impact also on the way cancers could be treated as classically cancer treatment strategies have been considered cancer type specific. The hundreds of different forms of cancers originating from different sites of organs and more specifically the different cell types are driven by a complex array of genomic alterations and no two cancers are identical. Thus, it is easy to argue that no two cancers respond similarly to a given therapy. How do we then match the patients with the right treatment? The pattern how specified genetic alterations in a given cellular lineage background result in tumors with different biology, pathology and treatment sensitivity are beginning to be characterized, and based on this information the patients could be classified further beyond the pathological and genetic features. In case of the novel immunotherapies, such tissue-agnostic cancer grouping and treatment strategy is already becoming a reality. In this concept identification of DNA mismatch repair defects confer into a biomarker that can be applied to patient stratification in multiple different cancer types. For other targeted therapeutics such as kinase inhibitors, the frequency of genomic aberrations in given types of cancers has defined the cancers with clinical use approval of the drugs, though the same oncogenic aberrations, even if rare, can be drivers also in other types of cancers. The ability to diagnostically confirm a patient’s tumor cells´ sensitivity towards targeted therapeutics following a genetic profiling could provide the needed evidence to use such therapeutics also to treat other rare cancers, “off-label”, and to confirm the patient specific efficacy before each treatment decision. The platforms for such ex vivo therapy efficacy screening exist, and clinical validation of these methods is underway.
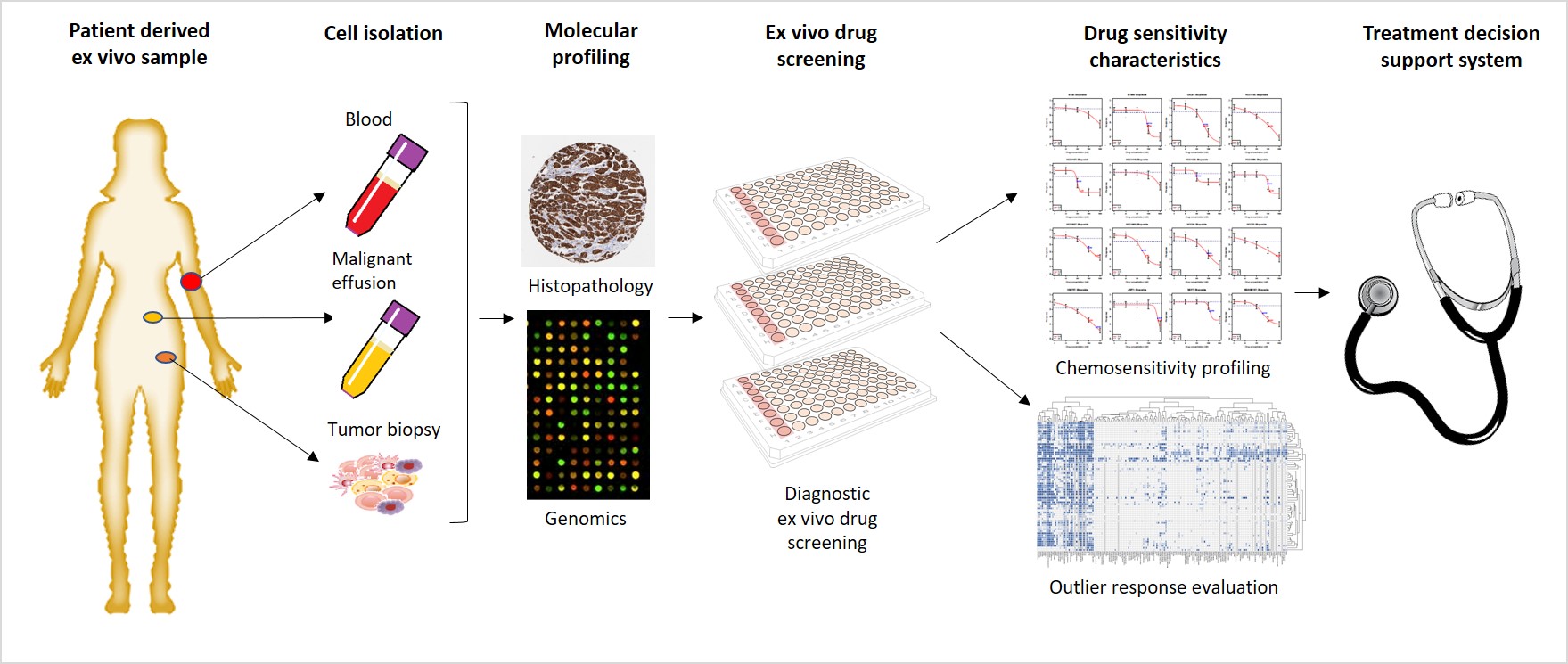
Tissue agnostic ex vivo evidence-based therapy
Genomic medicine offers a static view on the aberrations driving a cancer and suggests sensitivity towards given therapy, but it lacks on empirical evidence of efficacy for an individual patient. Due to the genetic complexity and lineage heterogeneity of cancer cells, the patient specific efficacy of therapies is unpredictable even within cancers of the same type. Missing this functional predictive aspect has hindered adaptation of genomic medicine to treatment of rare cancers. The ex vivo therapy efficacy screening strategy we are developing can be used for functional assessment of therapy efficacy in any solid cancer type. Currently the patients enrolled in the pre-clinical study represent more than 50 different solid cancers. The comparison of drug sensitivity of an individual patient’s tumor cells for 150 targeted cancer drugs across the developed pan-cancer drug sensitivity database allows assessment of patient and cancer specific drug responses. The outlier drug responses identified here indicate increased sensitivity of individual patient’s tumor towards specific drugs and highlight those that could result in a favourable clinical response. In combination with molecular pathology and genetic profiling of the patient’s tumor cells, the ex vivo drug sensitivity profiling can be used as empirical evidence to support treatment decisions for patients with a type of cancer for which there is no clinical evidence for that drug. The pan-cancer drug sensitivity comparison offers thus new insights in repurposing of targeted therapies and identification of commonalities of drug sensitivity patterns of rare cancers with the common types of cancers.
